Originally posted by xyyman:






quote:I'm not following that closely yet. If contamination were an element could it have possibly been from earlier periods of handling of the mummies?
Originally posted by Truthcentric:
[QB] The contamination charge is ridiculous. Most of the people working on Tut, Ramses III, and the rest would have either been European or modern Egyptian, not sub-Saharan African or Afro-Diaspora.
quote:keep in mind the OP cotains dialog from two different posters Noah and egypt1101 as shown
Originally posted by xyyman:
Hmmmmm! Let me see.......they expected the geographic origin(STR) to be European?
Only delusional Europeans would harbor such a thought. LOL!
As I said when the results were first published, before the DNATribes analyzed the data. I expected the result to be e1b1b, e1b1a, hg-A, hg-J in that order. Why? Geographic location.
But guess what? ***ALL EIGHT***of the Amarnas were sub-saharan. Euros are freaking out.
quote:Confusing to whom?
Originally posted by xyyman:
[QUOTE]Originally posted by the lioness,:
[qb] the term "Saharan-Arabian" has led to a lot of confusion


quote:Here are some usable tools.
Originally posted by Truthcentric:
The contamination charge is ridiculous. Most of the people working on Tut, Ramses III, and the rest would have either been European or modern Egyptian, not sub-Saharan African or Afro-Diaspora.
I tried using that DNA-Fingerprint program (I presume it was Amatch?) Noah mentioned in his last post, and so far I'm not even sure how to enter in the Amarna data. If you notice, each STR listed in the JAMA paper has two values (e.g. CSF1PO has 6 and 12) , but the Amatch program only gives you one text field per locus.
Additionally, does anyone know if DNA Tribes has Upper Egyptians in their database?
quote:Excuse me, but I recall Beyoku or Swenet were actually among the fiercest defenders of the DNATribes findings. I think you may be confusing their argument again.
Originally posted by Amun-Ra The Ultimate:
Even in this forum, some people like Swenet and Beyoku didn't like the results. Trying to say that Ancient Egyptian are closer match to some oasis in modern egypt than what the results show.
quote:Defending it and not liking the results (and try to make them say something else) are not the same thing!!
Originally posted by Truthcentric:
quote:Excuse me, but I recall Beyoku or Swenet were actually among the fiercest defenders of the DNATribes findings. I think you may be confusing their argument again.
Originally posted by Amun-Ra The Ultimate:
Even in this forum, some people like Swenet and Beyoku didn't like the results. Trying to say that Ancient Egyptian are closer match to some oasis in modern egypt than what the results show.
quote:And what would that be?
Originally posted by Amun-Ra The Ultimate:
Defending it and not liking the results (and try to make them say something else) are not the same thing!!
Since they know they can't attack the DNA Tribes results directly. They adopt a different strategy. They simply try to twist their meaning to make them means something else than what the results show directly.
quote:I knew you were going to ask that because it's a logical question. What are those "direct results" from DNA Tribes that I'm talking about?
And what would that be?
quote:Don't say that. Hawass surely has a lot of SSA on his team.
Originally posted by xyyman:
SSA researcher??? You kidding me! May be it was a family janitorial business...He! He!

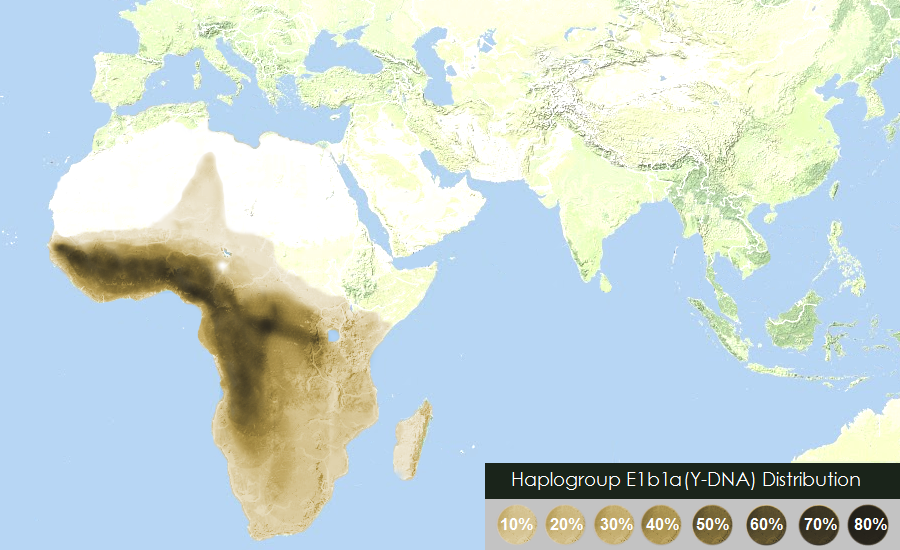
quote:^^^ This map is not that precise, not reflecting the Horn origin of the Hg
Originally posted by xyyman:

quote:
Originally posted by xyyman:


quote:--Alexandra Rosa et al.
Tajima's D and Fu's Fs, our unpublished data). An intriguing increased frequency of L0a1 in the Balanta might parallel A1-M31 and A3b2-M13 Y chromosomes in representing East African traces. Although the founder L0a1 haplotype is shared in an east-to-west corridor, the emerging lineages are exclusive of Guineans, indicating a rapid spread and local expansion after arrival.
The analysis of our data provides further evidence for the homogeneity of the Y chromosome gene pool of sub-Saharan West Africans, due to the high frequency of haplogroup E3a-M2. Its frequency and diversity in West Africa are among the highest found, suggesting an early local origin and expansion in the last 2030 ky. Hypothesizing on the existence of an important local agricultural centre, this could have supported a demographic expansion, on an E3a-M2 background, that almost erased the pre-existing Y chromosome diversity. Its pattern of diversity within Mandenka and Balanta hints at a more marked populational growth, these people possibly related to the local diffusion of agricultural expertise.
The Papel and Felupe-Djola people retain traces of their East African relatives, to which the short timescale of residence in Guinea-Bissau and higher isolation from major influences have contributed.
These minor imprints may represent movements from Sahel's more central and eastern parts, seen, for example, in the typically Ethiopian/Sudanese E3*-PN2 lineages that have reached Senegambia [2,3,5].
quote:Indeed. The only ones working in the labs of Egypt and handling the mummies are either Arabs or whites (European descent). LOL So the contamination theory is absurd.
Originally posted by Truthcentric:
The contamination charge is ridiculous. Most of the people working on Tut, Ramses III, and the rest would have either been European or modern Egyptian, not sub-Saharan African or Afro-Diaspora.
quote:Newsflash. The Egyptians WERE/ARE Northeast Africans! Egypt IS in Northeast Africa, the populations that were ancestral to Egyptians and became the Egyptians were northeast Africans. Exactly what evidence do have to the contrary?? Never mind, I don't want to see any of your pseudo-scholarly spam.
Originally posted by Clyde Winters:
It is sad that members of the Hamitic Union are trying to prove that the Egyptians are related to Northeast Africans as claimed by Egyptologist for the past 100 years. This proposition was never supported by the evidence,..
quote:The Meroitic Empire is also in Northeast Africa, twit. And there is no evidence of a mass exodus from Egypt to West Africa. The most pristine Egyptians can still be found today in the rural Nile Valley, and in the oases and deserts.
especially historical and archaeological evidence that indicated that after the fall of Egypt, Egyptians migrated into West Africa, and the Meroitic Empire not Northeast Africa.
![[Big Grin]](biggrin.gif)
quote:By itself that's a dumb argument. Yes, Ancient Egypt is where modern Egypt is now, but there's been a lot of population movements and changes in the genetic composition of the populations in Northeastern Africa since the last 6000 years. In fact, it's true for the rest of Africa and the world in general. Native America and Mesopotamia are other examples (see latest study about Mesopotamia). Same as the Bantu migration which changed the population composition all over Africa. It's always possible that some populations in modern Egypt we don't know about yet are closely related to Ancient Egyptians, but according to the genetic results thus far, they would need to be genetically closer to other African populations as a whole than to modern Eurasians or other modern North Africans. They will be as close to Ancient Egyptians as they are to other modern African populations.
Originally posted by Djehuti:
Newsflash. The Egyptians WERE/ARE Northeast Africans! Egypt IS in Northeast Africa
quote:
Originally posted by the lioness,:
quote:^^^ This map is not that precise, not reflecting the Horn origin of the Hg
Originally posted by xyyman:
.
.
quote:
Originally posted by xyyman:


quote:Stop bullshyting, how do the maps I posted of E1b1b1 and J1 originate in the Horn and Arabian penninsula respectively correspond to the Great lakes region?
Originally posted by xyyman:
I agree with you E and J1 is of Great Lakes/Sudan origin.
quote:Of course, and I never argued to the contrary. I actually forgot to mention in my last posting that the idiot Hamiticists are in denial that even though the Egyptians were northeast Africans, they and other northeast Africans still share genetic continuity with the rest of Africa including so-called 'Sub-Saharans' in West and Central Africa. As far as population movements are concerned, as I stated there is no evidence of Clyde's claims of a mass exodus to West Africa post-dynastic period. Instead what evidence we do have is Saharans during the Green Holocene moving the opposite direction i.e. from WEST to EAST. This is the reason why certain genetic elements like Benin HBS which is especially prominent in the oases areas.
Originally posted by Amun-Ra The Ultimate:
By itself that's a dumb argument. Yes, Ancient Egypt is where modern Egypt is now, but there's been a lot of population movements and changes in the genetic composition of the populations in Northeastern Africa since the last 6000 years. In fact, it's true for the rest of Africa and the world in general. Native America and Mesopotamia are other examples (see latest study about Mesopotamia). Same as the Bantu migration which changed the population composition all over Africa. It's always possible that some populations in modern Egypt we don't know about yet are closely related to Ancient Egyptians, but according to the genetic results thus far, they would need to be genetically closer to other African populations as a whole than to modern Eurasians or other modern North Africans. They will be as close to Ancient Egyptians as they are to other modern African populations.
quote:The difference is the spread of Islam when Arabs came into the region
Originally posted by xyyman:
Sudan, Ethiopia, Uganda ... What's the difference ?
On J1 , highest diversity is found in Sudan and Yemen. Ancestral form is found in Sudan and the Hadramat of Yemen. You know about that Yemeni group... right?
quote:Also Djehuti I would also like to expand on your point. I stated this on the Facebook.
Originally posted by Djehuti:
^ That's what I keep trying to tell Firewall. That even if J1 is of Southwest Asian origin, it still originated in tropical region among tropically adapted folks and NOT cold adapted modern 'white' Arab looking people.
quote:Of course, and I never argued to the contrary. I actually forgot to mention in my last posting that the idiot Hamiticists are in denial that even though the Egyptians were northeast Africans, they and other northeast Africans still share genetic continuity with the rest of Africa including so-called 'Sub-Saharans' in West and Central Africa. As far as population movements are concerned, as I stated there is no evidence of Clyde's claims of a mass exodus to West Africa post-dynastic period. Instead what evidence we do have is Saharans during the Green Holocene moving the opposite direction i.e. from WEST to EAST. This is the reason why certain genetic elements like Benin HBS which is especially prominent in the oases areas.
Originally posted by Amun-Ra The Ultimate:
By itself that's a dumb argument. Yes, Ancient Egypt is where modern Egypt is now, but there's been a lot of population movements and changes in the genetic composition of the populations in Northeastern Africa since the last 6000 years. In fact, it's true for the rest of Africa and the world in general. Native America and Mesopotamia are other examples (see latest study about Mesopotamia). Same as the Bantu migration which changed the population composition all over Africa. It's always possible that some populations in modern Egypt we don't know about yet are closely related to Ancient Egyptians, but according to the genetic results thus far, they would need to be genetically closer to other African populations as a whole than to modern Eurasians or other modern North Africans. They will be as close to Ancient Egyptians as they are to other modern African populations.
quote:that's bull
Originally posted by xyyman:
Like most genetic researchers I believe the Islamization was cultural and did not include movement of people.
quote:Yes, that's the impression I got as well. You see, there are many alleles present in the mummies that are not present in today's Egyptian population yet are found in other populations in Sub-Sahara. This points to common ancestry. However, note that Egypt in contrast to these other Sub-Saharan nations experienced dramatic population changes via incursions and immigrations.
Originally posted by Son of Ra:
Also Djehuti I would also like to expand on your point. I stated this on the Facebook.
Do you agree that the DNAtribes/JAMA results showed that the ANCESTORS of West Africans/Bantu people clustered with Ancient Egyptians, but that does not mean West Africans/Bantu people are the closet cluster group to the Ancient Egyptians? It just means the Ancient Egyptians and West Africans/Bantu people share a COMMON Ancestor and that West Africans/Bantu people do not descend from the Ancient Egyptians, but they do cluster with them due to both groups having a common ancestor. Again like you stated the ANCESTORS of West Africans/Bantu people originally lived in the Sahara when it was wet. And like you stated when it dried up, the groups migrated to two different regions. Remember E1b1a originated in East Africa in the first place. And I believe this video agrees with your points.
http://www.youtube.com/watch?v=98viuKQnIWU
So is this what DNAtribes is saying? Not that West Africans/Bantus are the closest cluster group to the AE or that they are descendants of the AE, but that they share a COMMON ANCESTOR with the AE.
quote:the above is from
Originally posted by xyyman:
From:
J1-M267 Y lineage marks climate-driven pre-historical human displacements Sergio Tofanelli,2009
To better understand the modes and timing of J1 dispersals, we reconstructed the genealogical relationships among 282 M267*G chromosomes from 29 populations typed at 20 YSTRs and 6 SNPs. Phylogenetic analyses depicted a new genetic background consistent with climate-driven demographic dynamics occurring during two key phases of human pre-history: (1) the spatial expansion of hunter gatherers in response to the end of the late Pleistocene cooling phases and (2) the displacement of groups of foragers/herders following the mid-Holocene rainfall retreats across the ***SAHARA AND ARABIA(part of greater Africa)****. Furthermore, J1 STR motifs previously used to trace Arab or Jewish ancestries were shown unsuitable as diagnostic markers for ethnicity
b] [/QB]
quote:I agree that the largest part of West African ancestry doesn't derived from Ancient Egyptians. West Africans are not direct descendant of Ancient Egyptians, at least not to a significant level. But both Western Africans, as well as other so-called "sub-Saharan" Africans, and Ancient Egyptians share common ancestors. Common ancestors postdating, for example, the OOA migration of non-Africans populations (but predating the birth of the Ancient Egyptian state).
Originally posted by Djehuti:
^ That's what I keep trying to tell Firewall. That even if J1 is of Southwest Asian origin, it still originated in tropical region among tropically adapted folks and NOT cold adapted modern 'white' Arab looking people.
quote:Of course, and I never argued to the contrary. I actually forgot to mention in my last posting that the idiot Hamiticists are in denial that even though the Egyptians were northeast Africans, they and other northeast Africans still share genetic continuity with the rest of Africa including so-called 'Sub-Saharans' in West and Central Africa. As far as population movements are concerned, as I stated there is no evidence of Clyde's claims of a mass exodus to West Africa post-dynastic period. Instead what evidence we do have is Saharans during the Green Holocene moving the opposite direction i.e. from WEST to EAST. This is the reason why certain genetic elements like Benin HBS which is especially prominent in the oases areas.
Originally posted by Amun-Ra The Ultimate:
By itself that's a dumb argument. Yes, Ancient Egypt is where modern Egypt is now, but there's been a lot of population movements and changes in the genetic composition of the populations in Northeastern Africa since the last 6000 years. In fact, it's true for the rest of Africa and the world in general. Native America and Mesopotamia are other examples (see latest study about Mesopotamia). Same as the Bantu migration which changed the population composition all over Africa. It's always possible that some populations in modern Egypt we don't know about yet are closely related to Ancient Egyptians, but according to the genetic results thus far, they would need to be genetically closer to other African populations as a whole than to modern Eurasians or other modern North Africans. They will be as close to Ancient Egyptians as they are to other modern African populations.
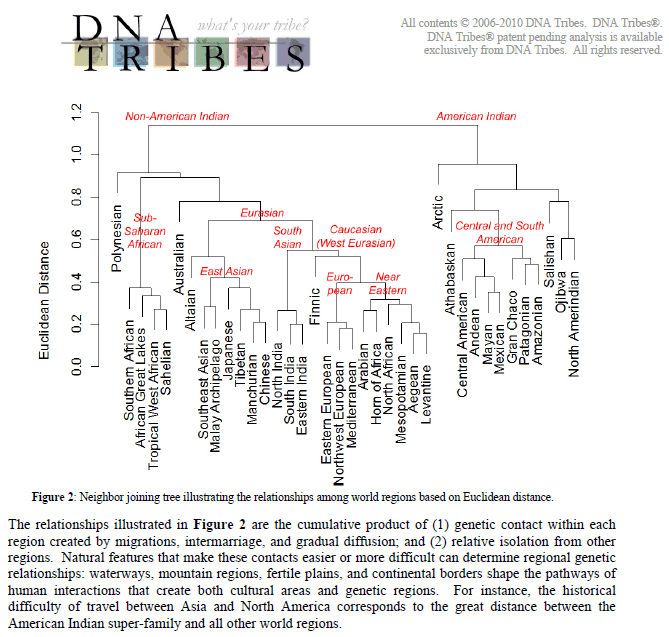
quote:You're mistaken. The DNA Tribes results, which used autosomal STR DNA, DOES say that Ancient Egyptian cluster more with modern Great Lakes Africans, Southern Africans and Western Africans population than any other population on earth. For example, Ancient Egyptians are genetically closer to populations in Western Africa than populations in modern Egypt.
Originally posted by Son of Ra:
So is this what DNAtribes is saying? Not that West Africans/Bantus are the closest cluster group to the AE or that they are descendants of the AE, but that they share a COMMON ANCESTOR with the AE.
quote:that article argues near eastern origin for J1, M43 and M78 that expanded from there into other parts of Arabian penninsula and across the Sahara
Originally posted by xyyman:
Nice ... get us back on track. At least we agree it had nothing to do with Islamic expansion. We can discuss its origin in another thread. Or just read my post on ESR .
quote:There must have been some Western Asian ancestry in Ancient Egypt, but it must have been minimal. Ancient Egyptians mummies don't cluster with Levantine or Western Asian populations. The Beyoku's preview study, while being an unproven source, gives the good idea. For example, in the Beyoku post we can see for example one J hg, among 21 mummies, but with an African mtDNA L hg on his mother side. So it was from a person who was admixed to some, probably low, degree.
Originally posted by xyyman:
Nice ... get us back on track. At least we agree it had nothing to do with Islamic expansion. We can discuss its origin in another thread. Or just read my post on ESR .
quote:I don't have to disprove something that you haven't proven and goes against common genetic knowledge.
Originally posted by xyyman:
I am going to be nice with you . Remember Beyoku met a truck head on and he was fatally wounded. So I am going to pose the question to you. Why isn't J1 African? Careful, don't stick you foot in your mouth.
quote:Common genetic knowledge doesn't say anything about Ancient Egyptians. For that we only got the JAMA and BMJ studies, including the fact that Ramses III is E1b1a/E-M2, and the DNA Tribes analysis of them, which I use to prove or disprove some points. There's not any genetic study that attributes the origin of J1 to Africa. I don't know what Mike has anything to do with anything, I practically never read his posts in my life, or at least don't remember them, but you must really like him to talk about him that way. You seem to like what he does. Are you his buddy or something? Is it another one of your ID here?
Originally posted by xyyman:
Common genetic knowledge? Do you know the AEians were Caucasoids?
Lucky I am not Mike. He likes to bitch slap the Negros when they screwup . But I am a nice guy.
quote:Of course that nonsense caucasoids claim is long overdue.
Originally posted by xyyman:
Listen babe...eh..dude. did you read Tofanelli paper, including the supplementals? Unlike most brothas here I believe many of the so-called "Caucasoid" North Africans are indigenous. No admixture needed. I am on the fence on J1 origins. Like most genetic researchers I believe the Islamization was cultural and did not include movement of people. J2 was introduced may be during Ottoman period that is why it is localized in the cities while J1 is more widespread indicating a prehistorical spread. I need to do a refresher on Tofanelli.
I have an entire spread on J on ESR.
This is one of the few times DJ is not sucking up to someone and has something worthwhile to add. Tropical Black Yemini and Sudanese ..whats the difference?
quote:--Meredith F. Small*
Morphological variation of the skeletal remains of ancient Nubia has been traditionally explained as a product of multiple migrations into the Nile Valley. In contrast, various researchers have noted a continuity in craniofacial variation from Mesolithic through Neolithic times. This apparent continuity could be explained by in situ cultural evolution producing shifts in selective pressures which may act on teeth, the facial complex, and the cranial vault.
A series of 13 Mesolithic skulls from Wadi Halfa, Sudan, are compared to Nubian Neolithic remains by means of extended canonical analysis. Results support recent research which suggests consistent trends of facial reduction and cranial vault expansion from Mesolithic through Neolithic times.
quote:This is from a older thread.
Originally posted by xyyman:
Last off topic comment. @ Lioness . I just had an Eureka!! moment. Looking at map you just posted I just realize that J1, E1b1b and E1b1a have similar geographic spread. Reading through the Tafonelli paper I see you have a relevant section highlighted. I saw that too. Which makes me lean futher to an African origin of J1. In the Great Lakes region. I need to follow up on the diversity. The paper does not have the raw data. It just has a chart in the supplementals. I was curios why they lump Ethiopians in with Eurasuans but Qatar , Tunisia , Iraq etc in with Arabic
Agreed the paper points to not an African origin. But it points to a spread PRIOR to the Islamic Expansion. As I said to be continued.
One paper I cited on ESR said eastern origin but when you look at the raw data Sudanese and Yemen diversity was equivalent. That is why I look at the raw data. The jury is still out but I am leaning to an African origin for J1.

quote:Blame it on the Jews,
Originally posted by xyyman:
@ Ultimate. What do you make of this? What is the story here? I am trying to pick out a trend. This if from Tofanelli latest paper on J1. I need some help here.
Hint: what is odd here is the highest diversity is found in....Portugal. does that mean J1 is Portugese in origin? Help me out here.

quote:I beg your pardon??! I took me a while to realize that this post was addressed to me. I mean "Hindu ass"?!
Originally posted by xyyman:
That is why you always come across a kiss ass. You really have a bad habit of not reading an article but insist on getting your Hindu ass to butt in.
![[Eek!]](eek.gif) LMAO
LMAO ![[Big Grin]](biggrin.gif) Exactly what makes you think I'm Hindu?! Also, I can read perfectly fine unlike YOU! Your dumb ass has the bad habit of reading an article without fully understanding it, and then putting coming to your own erroneous conclusions based misinterpreting what was truly said!!
Exactly what makes you think I'm Hindu?! Also, I can read perfectly fine unlike YOU! Your dumb ass has the bad habit of reading an article without fully understanding it, and then putting coming to your own erroneous conclusions based misinterpreting what was truly said!! quote:As I've always stated, the source of the expansion is ARABIA. Whether you include Arabia as part of Africa or Asia, is up to you. Also, while many the J1 subclades in Africa ARE due to Islamic expansion, not ALL of them are!! I already stated that there are J1 hgs present in the Horn region that predate Proto-Semitic, let alone the Arab language and Islam!
Quote by DJ: "Much of the J lineages in Africa, is the result of Islamic invasion.
Quote by Tofanelli: To better understand the modes and timing of J1 dispersals, we reconstructed the genealogical relationships among 282 M267*G chromosomes from 29 populations typed at 20 YSTRs and 6 SNPs. Phylogenetic analyses depicted a new genetic background consistent with climate-driven demographic dynamics occurring during two key phases of human pre-history: (1) the spatial expansion of hunter gatherers in response to the end of the late Pleistocene cooling phases and (2) the displacement of groups of foragers/herders following the mid-Holocene rainfall retreats across the SAHARA AND **ARABIA**(part of greater Africa). Furthermore, J1 STR motifs previously used to trace Arab or Jewish ancestries were shown unsuitable as diagnostic markers for ethnicity
Do you understand what he just said there DJ? I will help you. J1 had NADA, Nothing to do with Islamic Expansion.
![[Embarrassed]](redface.gif)
quote:Correct! And this explains the STRs and SNP results of DNA Tribes. I have been repeating this for years well before the results.
Originally posted by Amun-Ra The Ultimate:
I agree that the largest part of West African ancestry doesn't derived from Ancient Egyptians. West Africans are not direct descendant of Ancient Egyptians, at least not to a significant level. But both Western Africans, as well as other so-called "sub-Saharan" Africans, and Ancient Egyptians share common ancestors. Common ancestors postdating, for example, the OOA migration of non-Africans populations (but predating the birth of the Ancient Egyptian state).
quote:True. I never denied that there were migrations west during the Green Sahara. I merely disavowed that the claim of post-dynastic migrations. That there were migrations east to west by common Saharan ancestors explains many striking similarities like the Bes figure, certain staff figures like was, ritual statuary, votives, etc. etc.
I don't agree with you that the direction of genetic exchange within the Green Sahara populations, which include the Nile and the future location of Ancient Egypt, was only west to east. It was in both directions. West to east and East to West. The Green Sahara period, with its up and down, was a relatively long period in human history like at least 3000 years. We're in the years 2000s, and we talk about 3000 years here, so there has been time for a lot of population movements and admixture in all directions during the Green Sahara period.
quote:Yes, and really there was no "Sahara" or at least the desert was confined to smaller areas of the north. This is why the attempt of Euronuts to divide and segregate Africans is hilarious. And that includes the Hamitonuts!
During the Green Sahara period, predating the formation of the Ancient Egyptian state, "sub-Sahara" Africa so to say, was shifted northward. So rain, vegetation, animals and humans populations that followed them moved from "sub-Sahara" Africa to Northeastern Africa and the Sahara in general for at least 3000 years.
quote:Definitely Afrisian originated in northeast Africa in relatively more recent time compared to other African language phyla as that is its area of greatest diversity. And while Niger-Congo is greatest concentrated in West and Central Africa and Nilo-Saharan largely in East Africa, there is linguistic evidence to suggest that both phyla share a common ancestry. And that a remote branch of Niger-Congo (Kordofan) is still spoken in Sudan.
Also a bit before that time, or at the edge of it, all modern African genetic groups (like E-P2, probably true to for downstream A and B haplogroups too) and modern African languages phyla originate around northeastern Africa (around Sudan, Southern Egypt, Ethiopia, Somalia, etc). So most of all the modern Africans ancestry, as well as the Ancient Egyptian ancestry, can be found in eastern/northeastern Africa. It's afterward that they migrated to the rest of Africa like what would become the Green Sahara (probably meeting small group of humans there). Then when the Sahara got dry again, they had to move to Western Africa, Southern Africa, the Nile, etc (again meeting very small group of humans there. A00, Mbuti, etc). All those small groups of humans were absorbed by the migration of a much more demographically large African population from the Sahara. Leaving almost no trace of them in Western Africa (although the recently discovered A00 hg, is probably small traces of them).
quote:This actually debunks your theory, because according to your theory; Hg J1 should have arisen in Southern Portugal first.
Originally posted by the lioness,:
quote:Blame it on the Jews,
Originally posted by xyyman:
@ Ultimate. What do you make of this? What is the story here? I am trying to pick out a trend. This if from Tofanelli latest paper on J1. I need some help here.
Hint: what is odd here is the highest diversity is found in....Portugal. does that mean J1 is Portugese in origin? Help me out here.

notice on the chart
Is-Ash, the red diamond is also fairly high on the chart (Ashkenazi from Israel)
It is probably Jews in Portugal, I also see Omanese nearby
Am J Phys Anthropol. 2010 Mar;141(3):373-81. doi: 10.1002/ajpa.21154.
Phylogeographic analysis of paternal lineages in NE Portuguese Jewish communities.
Nogueiro I, Manco L, Gomes V, Amorim A, Gusmão L.
Source
Departamento de Antropologia, Universidade de Coimbra, 3000 Coimbra, Portugal.
Abstract
The establishment of Jewish communities in the territory of contemporary Portugal is archaeologically documented since the 3rd century CE, but their settlement in Trás-os-Montes (NE Portugal) has not been proved before the 12th century. The Decree of Expulsion followed by the establishment of the Inquisition, both around the beginning of the 16th century, accounted for a significant exodus, as well as the establishment of crypto-Jewish communities. Previous Y chromosome studies have shown that different Jewish communities share a common origin in the Near East, although they can be quite heterogeneous as a consequence of genetic drift and different levels of admixture with their respective host populations. To characterize the genetic composition of the Portuguese Jewish communities from Trás-os-Montes, we have examined 57 unrelated Jewish males, with a high-resolution Y-chromosome typing strategy, comprising 16 STRs and 23 SNPs. A high lineage diversity was found, at both haplotype and haplogroup levels (98.74 and 82.83%, respectively), demonstrating the absence of either strong drift or founder effects. A deeper and more detailed investigation is required to clarify how these communities avoided the expected inbreeding caused by over four centuries of religious repression. Concerning haplogroup lineages, we detected some admixture with the Western European non-Jewish populations (R1b1b2-M269, approximately 28%), along with a strong ancestral component reflecting their origin in the Middle East [J1(xJ1a-M267), approximately 12%; J2-M172, approximately 25%; T-M70, approximately 16%] and in consequence Trás-os-Montes Jews were found to be more closely related with other Jewish groups, rather than with the Portuguese non-Jewish population.
_________________________________________
xyyman always compare the gentics with history:
The history of the Jews in Portugal reaches back over two thousand years and is directly related to Sephardi history, a Jewish ethnic division that represents communities who have originated in the Iberian Peninsula (Portugal and Spain).
Jewish populations have existed on the area even before the country was established, back to the Roman era, or even before an attested Jewish presence in Portuguese territory, however, can only be documented since 482 CE.[1] With the fall of the Roman Empire, Jews were persecuted by the Visigoths and other European Christian kingdoms which controlled the area after that period.
In 711, the Moorish invasion of the Iberian Peninsula was seen by the many in the Jewish population as a liberation, and marked as the beginning of what many have seen as the Golden age of Jewish culture in the Iberian Peninsula (the Islamic Al-Andalus), even if the Jews, as well as the Christians (the Mozarabs of the Visigothic rite), under Muslim rule were considered Dhimmi, and had to pay a special tax.
Rapidly in the 8th century, the Christian kingdoms of the north mountainous areas of the Iberian Peninsula (Kingdom of Asturias) started a long military campaign against the Muslim invaders, the Reconquista. The Jews, since many knew the Arabic language, were used by the Christians as both spies and diplomats on this campaign that took centuries. This granted them some respect, although there was always prejudice.
___________________________________________
Landing near Algeciras in the spring of 711, the Muslim Moors (mainly Berbers with some Arabs) from North Africa invaded the Iberian peninsula,
Conversos were subject to suspicion and harassment from both the community they were leaving and that which they were joining. Both Christians and Jews called them tornadizo (renegade). James I, Alfonso X and John I passed laws forbidding the use of this epithet. This was part of a larger pattern of royal protection, as laws were promulgated to protect their property, forbid attempts to reconvert them, and regulate the behavior of the conversos themselves, preventing their cohabitation or even dining with Jews, lest they convert back.
The conversos did not enjoy legal equality. Alfonso VII prohibited the "recently converted" from holding office in Toledo. They had both supporters and bitter opponents within the Christian secular of general acceptance, yet they became targets of occasional pogroms during times of extreme social tension (as during an epidemic and after an earthquake). They were subject to the Spanish and Portuguese inquisitions.
While pure blood (so-called limpieza de sangre) would come to be placed at a premium, particularly among the nobility, in a 15th-century defense of conversos, Bishop Lope de Barrientos would list what Roth calls "a veritable 'Who's Who' of Spanish nobility" as having converso members or being of converso descent. He pointed out that given the near-universal conversion of Iberian Jews during Visigothic times, (quoting Roth) "[W]ho among the Christians of Spain could be certain that he is not a descendant of those conversos?"
According to a widely publicised study (December 2008) published in the American Journal of Human Genetics, 19.8 percent of modern Spaniards (and Portuguese) have DNA reflecting Sephardic Jewish ancestry (compared to 10.6 percent having DNA reflecting Moorish ancestors).[1] The Sephardic result is in contradiction [2][3][4] or not replicated in all the body of genetic studies done in Iberia and has been relativized by the authors themselves[1][5][6][7] and questioned by Stephen Oppenheimer, who estimates that much earlier migrations, 5,000 to 10,000 years ago from the Eastern Mediterranean, might also have accounted for the Sephardic estimates. "They are really assuming that they are looking at this migration of Jewish immigrants, but the same lineages could have been introduced in the Neolithic".[8] The same authors in also a recent study (October 2008) attributed most of those same lineages in Iberia and the Balearic Islands as of Phoenician origin.[9] The rest of genetic studies done in Spain estimate the Moorish contribution ranging from 2.5/3.4%[10] to 7.7%.[11]
quote:Quote me where I gave such a theory
Originally posted by Troll Patrol aka Ish Gebor:
This actually debunks your theory, because according to your theory; Hg J1 should have arisen in Southern Portugal first.
quote:That's exactly what I'm saying.
Originally posted by Troll Patrol aka Ish Gebor:
The thread and study by Beyoku, supports a probable East origin because of specific basic locus and alleles.
And yes, Sephardim Portuguese jews are eventually from the "Middle-East". So the high resolution fits perfectly. [/QB]
quote:Hmmmm,
Originally posted by the lioness,:
quote:Quote me where I gave such a theory
Originally posted by Troll Patrol aka Ish Gebor:
This actually debunks your theory, because according to your theory; Hg J1 should have arisen in Southern Portugal first.
I have no theory that hg J1 arose in Portugal. That was something xyyman asked about.
Origin is usually suggested by both frequency and diversity, I stated it earlier and put up a frequency chart that shows frequency highest in the Caucus and Arabia.
xyyman put up a chart that seems to show Portugese with some of the highest diversity. I don't know the exact context.
I put up information explaining recent CE reasons for the diversity not origin.
quote:That's exactly what I'm saying.
Originally posted by Troll Patrol aka Ish Gebor:
The thread and study by Beyoku, supports a probable East origin because of specific basic locus and alleles.
And yes, Sephardim Portuguese jews are eventually from the "Middle-East". So the high resolution fits perfectly.
However xxyman thinks J1 originates in Africa na he stated it on the pervious page.
He posted this chart on diversity to try to suggest that J1 coudn't have orignated in Portugal so therefore diversity is irrelevant.
Diversity is not irrelevant but it doesn't explain orign in this case of Portugal on the chart he posted.
You have not been following well. I stated several times J1 is near eastern.
xyyman's postion is that it is Sudanese origin [/QB]


quote:--Chiara Batini et al.
Sub-Saharan African Y chromosome diversity is represented by five main haplogroups (hgs): A, B, E, J, and R (Underhill et al. 2001; Cruciani et al. 2002; Tishkoff et al. 2007).
Hgs J and R are geographically restricted to eastern and central Africa, respectively, whereas hg E shows a wider continental distribution (see also Berniell-Lee et al. 2009; Cruciani et al. 2010).
[...]
All the samples included here were genotyped for ten STRs: DYS19, DYS389-I, DYS389-II (the allele reported in supplementary table S1, Supplementary Material online, has been obtained by subtracting the DYS389-I allele), DYS390, DYS391, DYS392, DYS393, DYS437, DYS438, and DYS439. A subset of the samples was tested for an additional five loci (DYS448, DYS456, DYS458 , DYS635, and Y-GATA-H4). In the statistical analyses, specific loci (DYS385, DYS389-II, DYS390, DYS448, and DYS635) were excluded due to allelic homoplasy as reported in the NIST Y-STR Fact Sheets (see Web resources in Acknowledgments). Following this, eight STR loci were used in both phylogeographic and intralineage analyses in order to maintain broad population coverage.
[...]
The starting set of markers comprised the 8 STRs used for Network analysis and diversity estimates and was extended to 11 by including DYS456, DYS458 , and YGATA-H4 loci. Due to multistep correction, different sets of STRs were used (supplementary table S7, Supplementary Material online), and the average mutation rate was estimated using locus-specific values (YHRD, release33; Willuweit et al. 2007).
quote:http://hamiticunion.proboards.com/index.cgi?board=general&action=display&thread=41
At the encouragement of another HU member, I have decided to start an open access library of notable works on or with direct relevance to Hamitic history, culture and biology. It is meant to complement the Hamitic Physical Anthropology thread by linking directly to complete materials on the same subject as opposed to excerpts from select chapters. Some of the more prominent examples of such classic works can be downloaded below, with more to follow
quote:In the meanwhile,
Every genetic study extant places the origins of haplogroup E either in Northeast Africa or in Eurasia. None associate the clade's origins directly with Negroid populations. Despite this, confusion (primarily among laypeople) seems to surround the haplogroup's population affinities. This mainly has to do with the fact that numerous (though not all) modern Negroid peoples carry the clade. This thread helps explain why they do, as well as how, when and through what mechanism(s) those Negroid peoples acquired haplogroup E in the first place from the Hamitic folk it originated with.
Read more: http://hamiticunion.proboards.com/index.cgi?board=general&action=display&thread=4#ixzz2mPl3u0Zn
quote:--Y-DNA Haplogroup Tree 2013
The DE haplogroup appeared approximately 50,000 years bp in North East Africa and subsequently split into haplogroup E that spread to Europe and Africa and haplogroup D that rapidly spread along the coastline of India and Asia to North Asia.
quote:--Y-DNA Haplogroup A and its Subclades - 2013
A1a-M31 is observed in northwestern Africans; A1b1a-M14 is seen among click language-speaking Khoisan populations. A1b1b-M32 has a wide distribution including Khoisan speaking and East African populations, and scattered members on the Arabian Peninsula
quote:--Y-DNA Haplogroup B and its Subclades - 2013
Y-DNA haplogroup B, like Y-DNA haplogroup A, is seen only in Africa and is scattered widely, but thinly across the continent. B is thought to have arisen approximately 50,000 years ago. These haplogroups have higher frequencies among hunter-gather groups in Ethiopia and Sudan, and are also seen among click language-speaking populations. The patchy, widespread distribution of these haplogroups may mean that they are remnants of ancient lineages that once had a much wider range but have been largely displaced by more recent population events.
Some geographic structuring is seen between the sub-groups B2a (B-M150) and B2b (B-M112). Sub-group B2b is seen among Central African Pygmies and South African Khoisan. Sub-group B2a is seen among Cameroonians, East Africans, and among South African Bantu speakers. B2a1a (B-M109) is the most commonly seen sub-group of B2a. About 2.3% of African-Americans belong to haplogroup B - with 1.5% of them belonging to the sub-group B2a1a.
quote:
Originally posted by the lioness,:
quote:that's bull
Originally posted by xyyman:
Like most genetic researchers I believe the Islamization was cultural and did not include movement of people.
quote:quote Tafonelli where he says Arab DNA had no effect on the genetics of Africa
Originally posted by xyyman:
You do realize Tafonelli agrees with me? The same paper you cited. Guess you did not read it before you posted.? That is not very smart.
quote:
Originally posted by the lioness,:
quote:that's bull
Originally posted by xyyman:
Like most genetic researchers I believe the Islamization was cultural and did not include movement of people.
quote:Thank you Mike for providing us with some humoristic moments on this forum. You show us that anybody can have an opinion which defies science and basic logic and hold on to it.
Originally posted by xyyman:
Would you give it up already Ultimate. There is very little "non-African" haplogroups. Aren't you reading the post ....or not understanding. R-V88 has recently been proven as NOT back-migration. M/N, female, is more diverse in Africa. Y- C, D, F etc is found all over Africa including West Africa would you be cognizant of the BS you are posting. It is a good idea to read other cited sources by other posters...it is called a well rounded education. That way you sound like you know what you are talking about, and not like a dogmatic idiot. Little advice.
quote:so you agree with Tofanelli who says that the first wave of J1 came from the Arabian penninusla into Africa in pre historic times?
Originally posted by xyyman:
Listen babe. I know you are not as dumb as the question you just asked. The title says it all . I quoted/bolded it in the abstract. Key words "climate driven" and "pre-history". I don't need to say more.
quote:
Originally posted by Amun-Ra The Ultimate:
The Y-DNA haplogroups situation is really simple:
_____________________________J M267
ABE=African
DCF=non-African
quote:Prior to Arab culture in Arab, the first civilization:
Originally posted by xyyman:
I agree with Tafonelli that it was NOT during Islamization. Where it originated ? .. I am on the fence. TP summed it up . My Eureka moment shows J1 mimics E1b1b. I need to follow up on J in the Lemba. Your J chart may need updating.
quote:
Originally posted by xyyman:
I said this so many times. I am hesitant to believe what white people write in their history books. White people lie. They falsify history, it is that simple. But based upon my experience they are less likely to falsify data. They will try to tell you what the data means. As an educated black people we must not fall for that trick. Got it Ultimate?! The mathematical model shows J1 spreading prior to the Islamic period which I agree with Tafonelli.That chart I put up would help resolve its origin. Portugal North, South and Central has high diversity but low frequency. J1 seems to mimic E1b1b. Ethiopia seems a practical point of origin. But I need more data.


quote:here is the Tofanelli 2009 he referred to contradicting you. If you were comprehending properly, in addition to J, he even refers to certain sub clades of E as near eastern
Originally posted by xyyman:
[QB] @ Lioness. I don't do we'll with hypotheticals. In 2009 study by Tofanelli posted by TP, he speculated J1 was an East African haplogroups. In this study we have a high frequency and high diversity in East Africa. Both E1b1b and J1 mimic each other . We know E is African .......
quote:The 2008 article is on a specific allelle an inconclusive
Originally posted by xyyman:
I guess you missed the ...."posted by TP. " It was Torafenelli earlier work.
Bottom line. Here are the facts:
East Africans has the highest frequency combined with highest diversity. Portuguese has a slightly higher diversity but substantial lower frequency. This applies to Black Sea area also. I am not sure what it means ....as yet. Ignoring a white man telling me what it means. The mathematical model suggest the foundation occurred around the Helocene
quote:Your little map and arrows are b.s. since its long been established that J1 entered the Horn during the Neolithic and its presence in southern Arabia was established before the mesolithic! Again the oldest subclades are found in Arabia NOT Anatolia or the Caucasus. And what about its ancestral clade un-derived J*? J* is found nowhere else but in the southern areas of Arabia and in the coastal Horn, its highest frequency is in Soqotra Island among the aborigines.
Originally posted by the lyinass,:
[/URL]
my arrows here for possible migration of J1
This a few thousand years before Islam, Mesopotamia marked purple was extending into the Arabian coast
The branch that went into Africa a bit later
quote:LOL Your post is nothing more than a prejudice generalization on whites. Not all white people are racist or have biased views. Yes racist bias exists but to blanket all whites of being guilty of this is a non-starter. Your post is also contradictory because you then cite and rely heavily on scientific studies conducted by and written by WHITES. It is this type of schizoid thinking that explains why you cannot comprehend writing that well or make these ridiculous assertions like me being 'Hindu'! LOL
Originally posted by xyyman:
I said this so many times. I am hesitant to believe what white people write in their history books. White people lie. They falsify history, it is that simple. But based upon my experience they are less likely to falsify data. They will try to tell you what the data means. As an educated black people we must not fall for that trick. Got it Ultimate?! The mathematical model shows J1 spreading prior to the Islamic period which I agree with Tafonelli.That chart I put up would help resolve its origin. Portugal North, South and Central has high diversity but low frequency. J1 seems to mimic E1b1b. Ethiopia seems a practical point of origin. But I need more data.
![[Big Grin]](biggrin.gif)
quote:
Originally posted by Djehuti:
quote:Your little map and arrows are b.s. since its long been established that J1 entered the Horn during the Neolithic and its presence in southern Arabia was established before the mesolithic! Again the oldest subclades are found in Arabia NOT Anatolia or the Caucasus. And what about its ancestral clade un-derived J*? J* is found nowhere else but in the southern areas of Arabia and in the coastal Horn, its highest frequency is in Soqotra Island among the aborigines.
Originally posted by the lyinass,:
[/URL]c
my arrows here for possible migration of J1
This a few thousand years before Islam, Mesopotamia marked purple was extending into the Arabian coast
The branch that went into Africa a bit later
quote:Actually the situation is a bit more complex. Haplogroup F , the ancestral clade of the vast majority of Eurasian lineages, has long been presumed to have originated in Asia. However there are significant traces of underived F* present in Africa, particularly among Sudanese tribes. Predictably, many experts then assumed that this was due to Arab genetic influence among these tribes. The problems with this hypothesis is that F is rare in Arabia with underived F* practically non-existent, while the same Sudanese tribes who carry F* show minimal frequencies of J while showing high frequencies of indigenous hgs like A and B. Furthermore, the recent genetic analysis of Neoltihic Nubian remains yield hg A as well as hg F*. All of this supports the theory that F* may have originated in Africa first before entering Southwest Asia and dispersing.
Originally posted by Amun-Ra The Ultimate:
The Y-DNA haplogroups situation is really simple:
ABE=African
DCF=non-African
Obviously since the main OOA migration of non-African there's been some level of admixtures between humans who stayed in Africa and humans who left Africa(F descendants mostly).
For example, some Cameroonians got the R haplogroup. R haplogroup in Africa is an example of the back migration of F descendants in Africa after many millennium of separation. R is an haplogroup descendant of the F haplogroup along with other haplogroup like R,H,I,J,K. Still in this case, it's most probable that if autosomal DNA were analyzed the full genome of those Cameroonians would be mostly African despite possessing a non-African haplogroup. Since Y-DNA and MtDNA haplogroups are only one line of descent. Same thing with ancient E or mtDNA L haplogroup found in Europe. In this case, even if some "native" Europeans got the E and mtDNA L haplogroup, before and around 10 000BC, their full genome is probably mostly Europeans.
The mtDNA situation is similarly simple.
L and possibly M1=African
non-L=non-African
quote:Even if the F and J haplogroup and almost all haplogroups in the world would be African (which is definitely false and ridiculous), it doesn't matter. Because F and J hg are rare among African people beside those sharing frontier with Eurasian populations.
Originally posted by Djehuti:
quote:Actually the situation is a bit more complex. Haplogroup F , the ancestral clade of the vast majority of Eurasian lineages, has long been presumed to have originated in Asia. However there are significant traces of underived F* present in Africa, particularly among Sudanese tribes. Predictably, many experts then assumed that this was due to Arab genetic influence among these tribes. The problems with this hypothesis is that F is rare in Arabia with underived F* practically non-existent, while the same Sudanese tribes who carry F* show minimal frequencies of J while showing high frequencies of indigenous hgs like A and B. Furthermore, the recent genetic analysis of Neoltihic Nubian remains yield hg A as well as hg F*. All of this supports the theory that F* may have originated in Africa first before entering Southwest Asia and dispersing.
Originally posted by Amun-Ra The Ultimate:
The Y-DNA haplogroups situation is really simple:
ABE=African
DCF=non-African
Obviously since the main OOA migration of non-African there's been some level of admixtures between humans who stayed in Africa and humans who left Africa(F descendants mostly).
For example, some Cameroonians got the R haplogroup. R haplogroup in Africa is an example of the back migration of F descendants in Africa after many millennium of separation. R is an haplogroup descendant of the F haplogroup along with other haplogroup like R,H,I,J,K. Still in this case, it's most probable that if autosomal DNA were analyzed the full genome of those Cameroonians would be mostly African despite possessing a non-African haplogroup. Since Y-DNA and MtDNA haplogroups are only one line of descent. Same thing with ancient E or mtDNA L haplogroup found in Europe. In this case, even if some "native" Europeans got the E and mtDNA L haplogroup, before and around 10 000BC, their full genome is probably mostly Europeans.
The mtDNA situation is similarly simple.
L and possibly M1=African
non-L=non-African
But I do agree that though Southwest Asia may be the launching pad for multiple OOA migrations, there was obviously some back-and-forth migrations between Southwest Asia and Africa, hence haplogroups G, I, J, K, R, and T in Africa. Just like Keita says, Paleolithic hunter-gatherers were constantly moving not only between 'sub-Sahara' and 'Supra-Sahara' but also between Supra-Sahara' and Eurasia, specifically Southwest Asia.

quote:I am posting from your favorite board. The ridiculing is way too funny.
Originally posted by the lioness,:
^^^^ as xyyman says, "overload"
the guy is spammng now. It's not cohaerant as dialog, just something to skip, 4 posts in a row talking to himself
quote:That's odd, I swear these sources say something else. How is that possible?
Originally posted by Amun-Ra The Ultimate:
The Y-DNA haplogroups situation is really simple:
ABE=African
DCF=non-African
Obviously since the main OOA migration of non-African there's been some level of admixtures between humans who stayed in Africa and humans who left Africa(F descendants mostly).
For example, some Cameroonians got the R haplogroup. R haplogroup in Africa is an example of the back migration of F descendants in Africa after many millennium of separation. R is an haplogroup descendant of the F haplogroup along with other haplogroup like R,H,I,J,K. Still in this case, it's most probable that if autosomal DNA were analyzed the full genome of those Cameroonians would be mostly African despite possessing a non-African haplogroup. Since Y-DNA and MtDNA haplogroups are only one line of descent. Same thing with ancient E or mtDNA L haplogroup found in Europe. In this case, even if some "native" Europeans got the E and mtDNA L haplogroup, before and around 10 000BC, their full genome is probably mostly Europeans.
The mtDNA situation is similarly simple.
L and possibly M1=African
non-L=non-African
quote:Y-DNA Haplogroup F and its Subclades - 2013
Y-DNA haplogroup F is the parent of all Y-DNA haplogroups G through T and contains more than 90% of the worlds population. Haplogroup F was in the original migration out of Africa, or else it was founded soon afterward, because F and its sub-haplogroups are primarily found outside, with very few inside, sub-Saharan Africa. The founder of F could have lived between 60,000 and 80,000 years ago, depending on the time of the out-of-Africa migration.
The major sub-groups of Haplogroup F are Haplogroups G, H, [IJ], and K, which are discussed elsewhere at this site. The minor sub-groups, F*, F1, and F2 have not been well studied, but apparently occur only infrequently and primarily in the Indian subcontinent. F* has been observed in two individuals in Portugal, possibly representing a remnant of 15th and 16th century contact of Portugal with India.
quote:http://www.familytreedna.com/public/albrosurnameproj/default.aspx?section=news
"I is a branch of haplogroup F* (M89 mutation), which first appeared in Africa some 45,000 years before the present. F* is believed to represent the "second-wave" of expansion out of Africa between 45 and 40 thousand years ago, that went directly to the Middle East"
quote:--University of Bridgeport (2011)
In human genetics, Haplogroup F* (M89) is a Y-chromosome haplogroup (Note: due to technical restrictions, the title of this page does not contain an "*").
This haplogroup first appeared in Africa some 45,000 years before present. It is believed to represent the "second-wave" of expansion out of Africa.
Haplogroup F* is an ancestral haplogroup to Y-chromosome haplogroups G (M201), H (M52), I (M170), J (12f2.1), and K (M9) along with its descendant haplogroups (L, M, N, O, P, Q, and R).
quote:That's odd, because here it does!
Originally posted by the lioness,:
quote:
Originally posted by Amun-Ra The Ultimate:
The Y-DNA haplogroups situation is really simple:
_____________________________J M267
ABE=African
DCF=non-African
quote:--Y-DNA Haplogroup Tree 2013
The DE haplogroup appeared approximately 50,000 years bp in North East Africa and subsequently split into haplogroup E that spread to Europe and Africa and haplogroup D that rapidly spread along the coastline of India and Asia to North Asia.
quote:Y-DNA Haplogroup Tree 2013, 3 October 2013
The BT haplogroup split from the root of the Y haplogroup tree 55,000 years before present (bp), probably in North East Africa. The CF(xDE) haplogroup was the common ancestor of all people who migrated outside of Africa until recent times. The defining mutation occurred 31-55,000 years bp in North East Africa and is still most common in Africa today in Ethiopia and Sudan. The DE haplogroup appeared approximately 50,000 years bp in North East Africa and subsequently split into haplogroup E that spread to Europe and Africa and haplogroup D that rapidly spread along the coastline of India and Asia to North Asia.
quote:Strange, from what I can recall, this is what they said?
Originally posted by Amun-Ra The Ultimate:
The Y-DNA haplogroups situation is really simple:
ABE=African
DCF=non-African
Obviously since the main OOA migration of non-African there's been some level of admixtures between humans who stayed in Africa and humans who left Africa(F descendants mostly).
For example, some Cameroonians got the R haplogroup. R haplogroup in Africa is an example of the back migration of F descendants in Africa after many millennium of separation. R is an haplogroup descendant of the F haplogroup along with other haplogroup like R,H,I,J,K. Still in this case, it's most probable that if autosomal DNA were analyzed the full genome of those Cameroonians would be mostly African despite possessing a non-African haplogroup. Since Y-DNA and MtDNA haplogroups are only one line of descent. Same thing with ancient E or mtDNA L haplogroup found in Europe. In this case, even if some "native" Europeans got the E and mtDNA L haplogroup, before and around 10 000BC, their full genome is probably mostly Europeans.
The mtDNA situation is similarly simple.
L and possibly M1=African
non-L=non-African
quote:--Chiara Batini et al.
Sub-Saharan African Y chromosome diversity is represented by five main haplogroups (hgs): A, B, E, J, and R (Underhill et al. 2001; Cruciani et al. 2002; Tishkoff et al. 2007).
Hgs J and R are geographically restricted to eastern and central Africa, respectively, whereas hg E shows a wider continental distribution (see also Berniell-Lee et al. 2009; Cruciani et al. 2010).
[...]
quote:--Fulvio Cruciani et al (2011)
The deepest branching separates A1b from a monophyletic clade whose members (A1a, A2, A3, B, C, and R) all share seven mutually reinforcing derived mutations (five transitions and two transversions, all at non-CpG sites). To retain the information from the reference MSY tree13 as much as possible, we named this clade A1a-T (Figure 1). Within A1a-T, the transversion V221 separates A1a from a monophyletic clade (called A2-T) consisting of three branches: A2, A3, and BT, the latter being supported by ten mutations (Figure 1).
quote:Can you explain why this is:
Originally posted by Amun-Ra The Ultimate:
The Y-DNA haplogroups situation is really simple:
ABE=African
DCF=non-African
Obviously since the main OOA migration of non-African there's been some level of admixtures between humans who stayed in Africa and humans who left Africa(F descendants mostly).
For example, some Cameroonians got the R haplogroup. R haplogroup in Africa is an example of the back migration of F descendants in Africa after many millennium of separation. R is an haplogroup descendant of the F haplogroup along with other haplogroup like R,H,I,J,K. Still in this case, it's most probable that if autosomal DNA were analyzed the full genome of those Cameroonians would be mostly African despite possessing a non-African haplogroup. Since Y-DNA and MtDNA haplogroups are only one line of descent. Same thing with ancient E or mtDNA L haplogroup found in Europe. In this case, even if some "native" Europeans got the E and mtDNA L haplogroup, before and around 10 000BC, their full genome is probably mostly Europeans.
The mtDNA situation is similarly simple.
L and possibly M1=African
non-L=non-African

quote:-- Semino et al.
The coding regions transitions are likely to change relatively slower than those of hypervariable segments, and hence, likely to remain intact within a clade. To assist in determining which clade to place a monophyletic unit, key coding region transitions have to be identified. In the case of M1, we were told:
We found 489C (Table 3) in all Indian and eastern-African haplogroup M mtDNAs analysed, but not in the non-M haplogroup controls, including 20 Africans representing all African main lineages (6 L1, 4 L2, 10 L3) and 11 Asians.
These findings, and the lack of positive evidence (given the RFLP status) that the 10400 C->T transition defining M has happened more than once, suggest that it has a single common origin, but do not resolve its geographic origin. Analysis of position 10873 (the MnlI RFLP) revealed that all the M molecules (eastern African, Asian and those sporadically found in our population surveys) were 10873C (Table 3). As for the non-M mtDNAs, the ancient L1 and the L2 African-specific lineages5, as well as most L3 African mtDNAs, also carry 10873C.
Conversely, all non-M mtDNAs of non-African origin analysed so far carry 10873T. These data indicate that the **transition 10400 C-->T, which defines haplogroup M**, arose on an African background characterized by the ancestral state 10873C, which is also present in four primate (common and pygmy chimps, gorilla and orangutan) mtDNA sequences.
quote:--Hans-Jürgen Bandelt et. 2006. EDS. Human Mitochondrial DNA and
"These indicate that the root of L3 gives rise to a multifurcation from a
single haplotype producing a number of distinct subclades... The
simplest explanation for this geographical distribution [haplogroups M
and N], however, is an expansion of the root type within East Africa,
where several independent L3 branches thrive, including a sister group
to L3, christened L4 (Kivisild et al. 2004; Chap. 7), followed by
divergence into haplogroups M and N somewhere between the Horn of
Africa and the Indian subcontinent. Since neither the L3 root type nor
any other descendants survive outside Africa, the root type itself must
have become extinct during a period of genetic drift in the founder
population as it diversified into haplogroups M and N, if the
diversification was outside Africa. If on the other hand the
diversification was indeed within East Africa, then Haplogroups M and
N must have either been carried out of Africa in their entirety or
subsequently have become extinct within Africa, with the singular
exception of the derived M1."
quote:--Paola Spinozzi, Alessandro Zironi .
Although Haplogroup M differentiated
soon after the out of Africa exit and it is
widely distributed in Asia (east Asia and
India) and Oceania, there is an
interesting exception for one of its more
than 40 sub-clades: M1.. Indeed this
lineage is mainly limited to the African
continent with peaks in the Horn of
Africa."
quote:-- Petraglia, M and Rose, J
..the M1 presence in the Arabian
peninsula signals a predominant East
African influence since the Neolithic
onwards.
quote:--SUVENDU MAJI, S. KRITHIKA and T. S. VASULU (2009)
Macrohaplogroup M (489-10400-14783-15043), excluding M1 which is east African, is distributed among most south, east and north Asians, Amerindians (containing a minority of north and central Amerindians and a majority of south Amerindians), and many central Asians and Melanesians.
quote:--Erwan Pennarun et al. Divorcing the Late Upper Palaeolithic demographic histories of mtDNA haplogroups M1 and U6 in Africa, BMC Evolutionary Biology 2012, 12:234
"No southwest Asian specific clades for M1 or U6 were discovered. U6 and M1 frequencies in North Africa, the Middle East and Europe do not follow similar patterns, and their sub-clade divisions do not appear to be compatible with their shared history reaching back to the Early Upper Palaeolithic."
quote:Go visit your favorite forum, hamiticcrap. Clown.
Originally posted by the lioness,:
^^^ redundant, obessive and desperate
![[Roll Eyes]](rolleyes.gif)
quote:--Marta Mirazón Lahr. 2005. The Evolution of Modern Human Diversity:
"others like Predomost and to a lesser degree Grimaldi and Teviec, are more prognathic like Skhul 5."
quote:--Chris Stringer, African Exodus ((Michael Witzel, The Origins of the World's Mythologies) 2013)
"...the Cro-Magnons, the presumed ancestors of modern Europeans....were more like present-day Australians or Africans..."
quote:--Stringer, C. B.(2003) Nature 423 , 692695. pmid:12802315
Today, most paleoanthropologists agree that the Cro-Magnons came from Africa (5).
quote:--B. Lewis et al. 2008. Understanding Humans: Introduction to Physical
"The so-called Old Man [Cro-Magnon 1] became the original model for
what was once termed the Cro-Magnon or Upper Paleolithic "race" of
Europe.. there's no such valid biological category, and Cro-Magnon 1 is
not typical of Upper Paleolithic western Europeans- and not even all that
similar to the other two make skulls found at the site. Most of the genetic
evidence, as well as the newest fossil evidence from Africa argue against
continuous local evolution producing modern groups directly from any
Eurasian pre-modern population.. there's no longer much debate that a
large genetic contribution from migrating early modern Africans infuenced
other groups throughout the Old World.
quote:--C. Loring Brace(2006)
If this analysis shows nothing else, it demonstrates that the oft-repeated European feeling that the Cro-Magnons are us (47) is more a product of anthropological folklore than the result of the metric data available from the skeletal remains.
quote:--[J Hum Evol 32 (1997a) 423], Bogin B, Rios L. et al.
It has been proposed that heat adapted, relatively long-legged Homo sapiens from Africa replaced the cold adapted, relatively short-legged Homo neandertalensis of the Levant and Europe
quote:--Trinkaus 2005
The subsequent post-28,000-B.P. Gravettian human sample of Europe includes numerous associated skeletons (Table 2) (Zilhão & Trinkaus 2002). Most of these specimens are fully modern in their morphology, and there is a persistence in them of both linear (equatorial) limb proportions and more "African" nasal morphology (Trinkaus 1981, Holliday 1997, Franciscus 2003). However, one Iberian specimen (Lagar Velho 1) exhibits Neandertal limb segment proportions and a series of relatively archaic cranial and postcranial features (Trinkaus & Zilhão 2002). In addition, central incisor shoveling, ubiquitous among the Neandertals, absent in the Qafzeh-Skhul sample, and variably present in the earlier European sample, persists at modest frequencies. And scapular axillary border dorsal sulci, an apparently Neandertal feature also absent in the Qafzeh-Skhul sample, is present
quote:-- Am J Phys Anthropol. 1975 May;42(3):351-69,
"Nor does the picture get any clearer when we move on to the Cro-Magnons, the presumed ancestors of modern Europeans. Some looked more like present-day Australians or Africans, judged by OBJECTIVE anatomical categorizations, as is the case with some early modern skulls from the Upper Cave at Zhoukoudian in China."
quote:--Encyclopedia of Human Evolution and Prehistory: Second Edition by Eric Delson
In modern humans, this elongation is a pattern characteristic of warm-adapted populations, and this physique may be an early Cro-Magnon retention from African ancestors. Similar retentions may be observed in certain indices of facial shape [ ...]
quote:I hate to do population structure with like 10 or so mummies. It's a number too small. But in general, current data seems to indicate, a bit like the Beyoku preview study, that Ancient Egyptians will be mostly from the A, B and E haplogroups. But there will also be some non-African haplogroups even at the formative years (neighboring regions always "leech" DNA from one another). So even among the first dynasty Ancient Egyptians, some R and J haplogroups, for example, will probably be present.
Originally posted by the lioness,:
.
which haplogroups of modern Egyptians are probably not ancient Egyptian haplogroups?
.
quote:M1 is more likely than possible East African in origin. That's what it says.
Originally posted by Amun-Ra The Ultimate:
Troll Patrol for one, your sources only confirm the graph I posted (with the OOA migration) and what I said about Y-DNA. As for mtDNA, it's easy to understand. L and M1 (possibly) = African, while non-L haplogroups present in African populations are an example of back migration of non-African people toward Africa and admixture with African people.
But more importantly by claiming hg rare in sub-Sahara Africa to be African in origin you play directly into the hand of Eurasian nuts who wants to disconnect Ancient Egypt from the rest of Africa (aka Sub-Sahara Africa). The Hamiticentric nut. Do you see that?
Of course, according to current DNA analysis of Ancient Egyptian mummies, Ancient Egyptians cluster with African populations ("sub-Saharan" Africans) not Eurasian populations, nor populations admixed with Eurasians to a significant degree. I think other mummies will also match Horn Africans too. Horn Africans which cluster with other 'Sub-Saharan Africans' on one of the DNA Tribes maps above. In general, I think the genetic distance between Ancient Egyptians and Africans, in general, will be lower than between Ancient Egyptians and non-Africans. Current genetic data seems to indicate that at the moment.
quote:In order to make such claim, you first need to sum which are present. Then look for the locus and alleles. By making such comparison you can draw lines or parallels with ancient once.
Originally posted by the lioness,:
.
Troll Patrol
which haplogroups of modern Egyptians are probably not ancient Egyptian haplogroups?
.
quote:I have answered you.
Originally posted by the lioness,:
I'm not making a claim I'm asking you Troll Patrol
which haplogroups of modern Egyptians are probably NOT ancient Egyptian haplogroups?
Now you're short on info ?????
quote:I didn't make a thesis. I asked you a question and you are afraid to answer it, lol
Originally posted by Troll Patrol aka Ish Gebor:
quote:I have answered you.
Originally posted by the lioness,:
I'm not making a claim I'm asking you Troll Patrol
which haplogroups of modern Egyptians are probably NOT ancient Egyptian haplogroups?
Now you're short on info ?????
In case you don't understand. It's your case, your thesis, you need to bring the evidence not me. Not the other way around.
quote:Because I can't answer a question which is unclear.lol
Originally posted by the lioness,:
quote:I didn't make a thesis. I asked you a question and you are afraid to answer it, lol
Originally posted by Troll Patrol aka Ish Gebor:
quote:I have answered you.
Originally posted by the lioness,:
I'm not making a claim I'm asking you Troll Patrol
which haplogroups of modern Egyptians are probably NOT ancient Egyptian haplogroups?
Now you're short on info ?????
In case you don't understand. It's your case, your thesis, you need to bring the evidence not me. Not the other way around.
quote:
Originally posted by the lioness,:
.
Troll Patrol
which haplogroups of modern Egyptians are probably not ancient Egyptian haplogroups?
.
code:The question now becomes, which one of the above hasn't been suggested as being African in origin?Old Kingdom (2686-2181 BCE)
yDna, mtDna
A-M13 L3f
A-M13 L0a1
B-M150 L3d
E-M2 L3e5
E-M2 L2a1
E-M123 L5a1
E-M35 R0a
E-M41 L2a1
E-M41 L1b1a
E-M75 M1
E-M78 L4b
J-M267 L3i
R-M173 L2
T-M184 L0a
Middle Kingdom (2055-1650 BCE)
A-M13 L3x
E-M75 L2a1
E-M78 L3e5
E-M78 M1a
E-M96 L4a
E-V6 L3
B-M112 L0b
quote:Yes, and I don't see how this contradicts anything I said. F-M89 was still present among Neolithic Nubians albeit at low frequencies with the predominant NRY being hg A. So what exactly what are you suggesting?.. that F-M89 and YAP (hg E) are both Eurasian as opposed to African in origin??
Originally posted by the lioness,:
http://etd2.uofk.edu/content/html/pdf/en/en.4312.pdf
Quote from: University of Khartoum
The area known today as Sudan may have been the scene of pivotal human evolutionary events, both as a corridor for ancient and modern migrations, as well as the venue of crucial past cultural evolution. Several questions pertaining to the pattern of succession of the different groups in early Sudan have been raised. To shed light on these aspects, ancient DNA (aDNA) and present DNA collection were made and studied using Y-chromosome markers for aDNA, and Y-chromosome and mtDNA markers for present DNA. Bone samples from different skeletal elements of burial sites from Neolithic, Meroitic, Post-Meroitic and Christian periods in Sudan were collected from Sudan National Museum. aDNA extraction was successful in 35 out of 76 samples, PCR was performed for sex determination using Amelogenin marker. Fourteen samples were females and 19 were males. To generate Y-chromosome specific haplogroups A-M13, B-M60, F-M89 and Y Alu Polymorphism (YAP) markers, which define the deep ancestral haplotypes in the phylogenetic tree of Y-chromosome were used. Haplogroups A-M13 was found at high frequencies among Neolithic samples. Haplogroup F-M89 and YAP appeared to be more frequent among Meroitic, Post-Meroitic and Christian periods. Haplogroup B-M60 was not observed in the sample analyzed.
quote:I never said that both F and J are African and I definitely never said all the other hgs I listed were African either. My original point is that it's likely that F-M89 for the reasons I stated as well as the study cited by Lioness.
Originally posted by Amun-Ra The Ultimate:
Even if the F and J haplogroup and almost all haplogroups in the world would be African (which is definitely false and ridiculous), it doesn't matter. Because F and J hg are rare among African people beside those sharing frontier with Eurasian populations.
quote:OK. No argument me. In fact I myself have been saying this all along! Perhaps you should try telling the idiot 'Hamiticists' that if you dare. It is they who try to separate African populations from one another in their debunked racial classifications of 'Hamite' vs. 'True Negro'.
So if Ancient Egyptian were only F and J haplogroups. It would disconnect Ancient Egyptians, current admixed East Africans and Eurasian North Africans from the rest of Africa. There would be African in geography only. Which is ridiculous.
So this would be another attempt to disconnect Ancient Egypt from the rest of Africa, thus the majority of African people.
When in reality, it's not the case. Ancient Egyptians according to the DNA Tribes results show that Africans are NOT genetically close to modern North Africans, modern Levantine, modern West Asian, modern admixed Horn Africans or modern Europeans. They are genetically closer to Great Lakes, Southern and Western Africans. Also BMJ identified Ramses III as being E1b1a/E-M2, an haplogroup mostly prevalent among so-called sub-Saharan Africans which form the genetic groups matching Ancient Egyptians.
Lets recall that Levantine, North Africans, Horn Africans, Great Lakes Africans, Tropical West Africans are genetic groupings of populations sharing the same genetic profile. Genetic families. Not ethnic, political or geographical groupings.
So that F and J hg being African would be just another attempt by horn supremacist as well as Eurasian supremacist (probably the same people) to disconnect North Africa (and Ancient Egypt) from the rest of Africa. Yes since Ancient Egyptian time, there's been a lot of foreign influx in those regions, but in ancient time the foreign influx from neighboring regions was present but much more limited.
Ancient Egyptians are NOT only Africans because of the geography. They also share biology, history, culture with other African people. When I say African people, I mean from North to South Africa. All African people from North to South Africa are close to each other genetically (beside coastal North Africans), culturally and historically. Due to 2 main things. 1)Shared origin (postdating the main OOA migration) 2)Admixtures
It's unfortunate that on the STR genetic distance tree by DNA Tribes (STRs used to match AE mummies), Horn Africans are not included in the Sub-Saharan Africans category, there was too much people of recent and ancient foreign origin in the STR samples of Horn Africans they took (you can see it in another DNA Tribes document). Similar genetic distance graph by DNA Tribes using SNP values, place back Horn Africans under the Sub-Saharan African fold.
This contrast with the other DNA Tribes Genetic Distance tree, using STR values, I posted above where the Horn Africans genetic group clustered more with the Eurasian genetic groups.
Nor Eurasian nor "admixed" Horn Africans match Ancient Egyptians but the genetic grouping called Sub-Saharan African by DNA Tribes (which btw could easily have been combined to form a bigger proportion thus a higher MLI scores. Mathematically it would render the denominator much smaller and the numerator bigger. Since MLI scores are proportions.).
So according to the DNA Tribes results, Ancient Egyptians matches so-called Sub-Saharan Africans to the great disarray of Eurasian nuts, admixed Horn Supremacists, coastal North African supremacists. It's not a great surprise because sub-Sahara Africa was shifted northward due to the monsoon rain during the Green Sahara period. Humans from the south followed rains, vegetations and animals northward. Seeking refuge afterward around at the Nile Valley, among other regions, after the dessication of the Green Sahara. Seeking greener pastures.
![[Embarrassed]](redface.gif)
quote:F-M89 is non-African. It's presence in Africa is a product of a back migration. Meroitic is like 300BC. The "Africanity" of modern Egyptian, and apparently Kushite, was already getting diluted at that time. The lioness quote just prove the introduction of foreign haplogroups in more recent times. In fact, even the Sudanese aDNA paper cited by lioness just above use F-M89 as a Eurasian markers. Here, I quote the document:
Originally posted by Djehuti:
quote:I never said that both F and J are African and I definitely never said all the other hgs I listed were African either. My original point is that it's likely that F-M89 for the reasons I stated as well as the study cited by Lioness.
Originally posted by Amun-Ra The Ultimate:
[qb]
Even if the F and J haplogroup and almost all haplogroups in the world would be African (which is definitely false and ridiculous), it doesn't matter. Because F and J hg are rare among African people beside those sharing frontier with Eurasian populations.
quote:So, according to the paper, Eurasian haplogroups are defined by F-M89.
Eurasian Haplogroups which are defined by
F-M89 against a background of haplogroup A-M13
quote:
Originally posted by the lioness,:

quote:So, what happend to people in Africa genetically before and after F-M89?
Originally posted by Amun-Ra The Ultimate:
quote:F-M89 is non-African. It's presence in Africa is a product of a back migration. Meroitic is like 300BC. The "Africanity" of modern Egyptian, and apparently Kushite, was already getting diluted at that time. The lioness quote just prove the introduction of foreign haplogroups in more recent times. In fact, even the Sudanese aDNA paper cited by lioness just above use F-M89 as a Eurasian markers. Here, I quote the document:
Originally posted by Djehuti:
quote:I never said that both F and J are African and I definitely never said all the other hgs I listed were African either. My original point is that it's likely that F-M89 for the reasons I stated as well as the study cited by Lioness.
Originally posted by Amun-Ra The Ultimate:
[qb]
Even if the F and J haplogroup and almost all haplogroups in the world would be African (which is definitely false and ridiculous), it doesn't matter. Because F and J hg are rare among African people beside those sharing frontier with Eurasian populations.
quote:So, according to the paper, Eurasian haplogroups are defined by F-M89.
Eurasian Haplogroups which are defined by
F-M89 against a background of haplogroup A-M13
quote:A loci is a specific mutation. Located in a gene. An allele is a variant of that specific gene.
Originally posted by the lioness,:
this is why you don't undertsand allele and loci, because the clade may not have the same geographic origin as the parent
like I have said if the hg is not African, what you and others do is follow branches back to the parent, they all go back to M168 and say that any Hg is therefore African
the "what was before" test etc
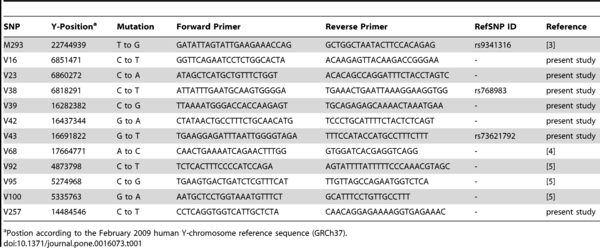
code:
SNP Location Haplogroup Mutations
M5 M C > T
M9 K, KR C > G
M11 L A > G
M45 P, PR G > A
M69 H T > C
M89 F, FR C > T
M96 E G > C
M122 O3 T > C
M168 CR C > T
M170 I A > C
M174 D T > C
M175 O T > A
M20 G G > T
M207 R A > G
M214 NO T > C
M304 J A > C
M343 R1b C > A
P36 Q G > T
SRY10831.1 BR A > G